Proteomics
Advertisement
Rover v.2.3.4
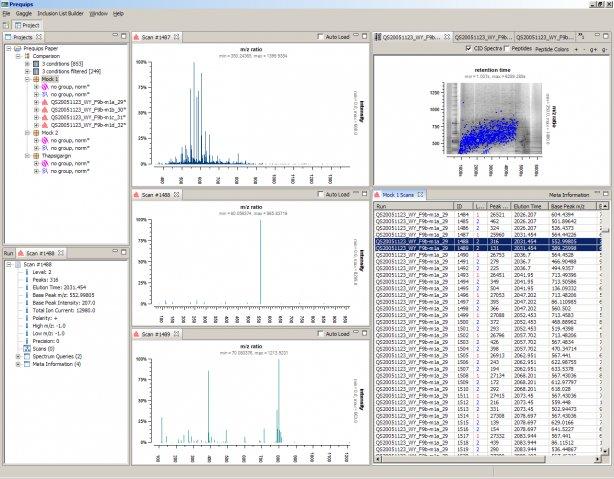
Rover is a software to visualize and validate quantitative proteomics data from different sources. Manual validation of regulated proteins in quantitative proteomics has always been a tedious process.
Advertisement
SPADE v.1.3
Build bioinformatics applications with this tool. The Structural Proteomics Application Development Environment (SPADE) is a free, Python-based, structural biology analysis and modeling toolkit,
ProteinID Finder v.1.8.2
A proteomics analyzation program capable of extracting information from the online universal protein resource database UniProt. ProteinID Finder is designed to simplify the database extraction and protein analyzation,
MzIdentML Viewer v.1.0
MzIdentML Viewer is a tiny, easy-to-use .MZID files viewer with just a click. This software makes use of the MzIdentML API. The Proteomics Information Group has developed MzIdentML,
EigenMS v.1.0
EigenMS is a stand-alone (or Matlab) bottom-up proteomics data normalization algorithms.
Maltcms v.1.2.0
Maltcms - Modular Application Toolkit for Chromatography Mass-Spectrometry is a JAVA API for preprocessing, alignment, analysis and visualization of data stored in open file formats used in Proteomics and Metabolomics research.
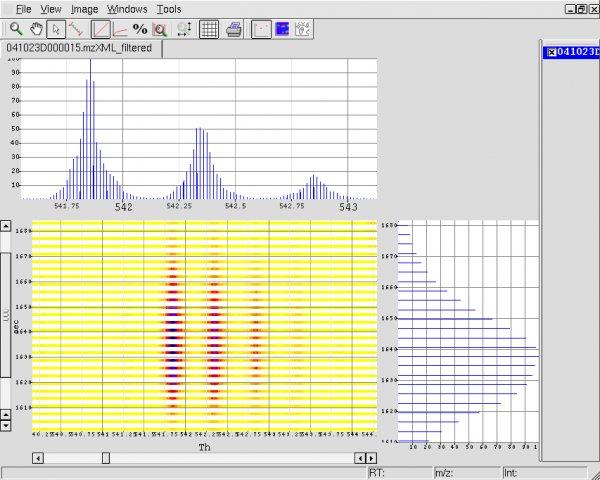
Open2Dprot v.1.0
Open2Dprot is an open-source proteomics project for the development of bioinformatic tools for n-dimensional protein expression data analysis of quantified protein expression across multiple samples from research experiments.
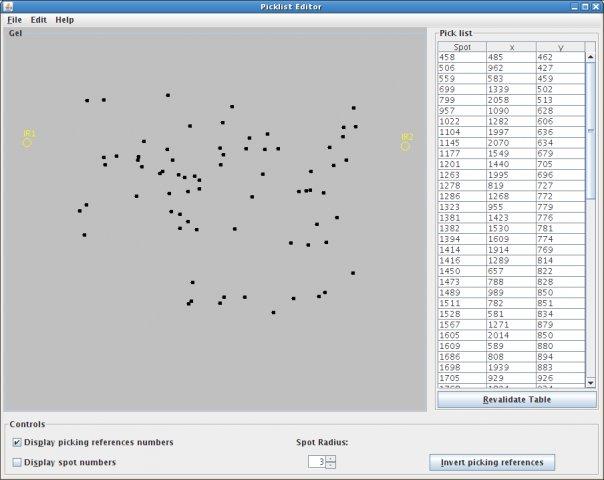
Pick List Editor v.1.0
The goal of Picklist Editor is to display and modify pick lists before spot picking (in proteomics).
Protein-ms v.1.0
Tools for mass spectrometry, especially for protein mass spectrometry and proteomics: Quantification tools, converters for Applied Biosystems (Q Star and Q Trap), calculation of in-silico fragmentation spectra, converter for Mascot result files